“Simpson’s paradox, or the Yule–Simpson effect, is a paradox in probability and statistics, in which a trend appears in different groups of data but disappears or reverses when these groups are combined.” ~ Wikipedia
I just got Simpson’s Paradox’ed and it feels so good! This is often a bad thing, as your analysis may come to a wrong conclusion if the data is viewed at the wrong level or you have latent grouping. But just now it helped me understand a problem and I figured I would tell the internet about it (like it or not).
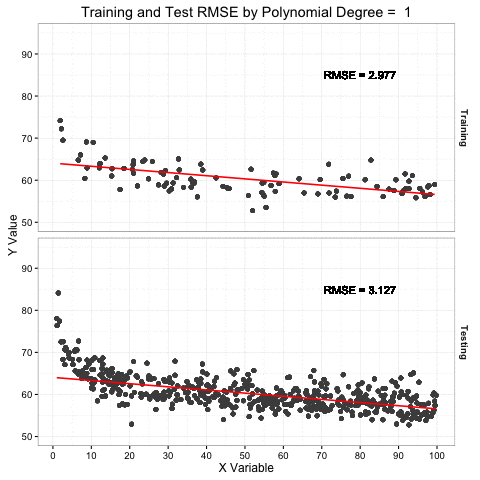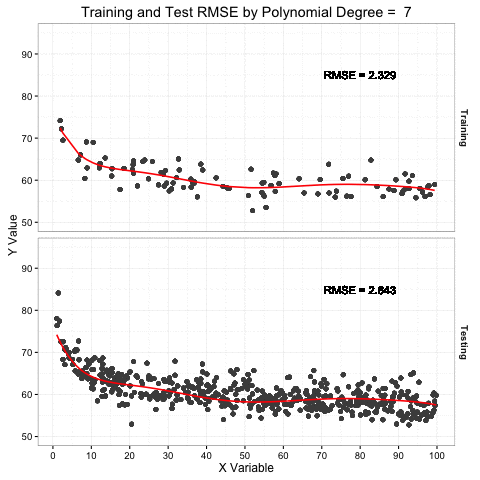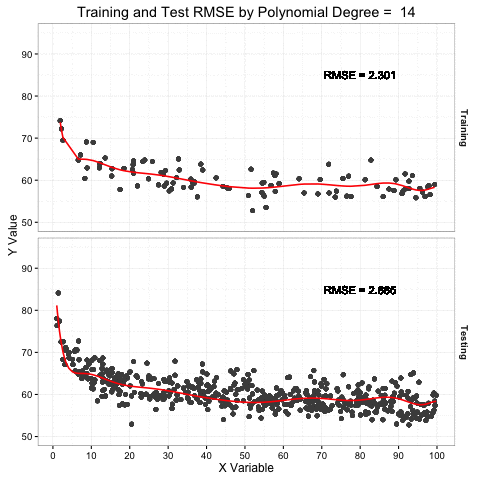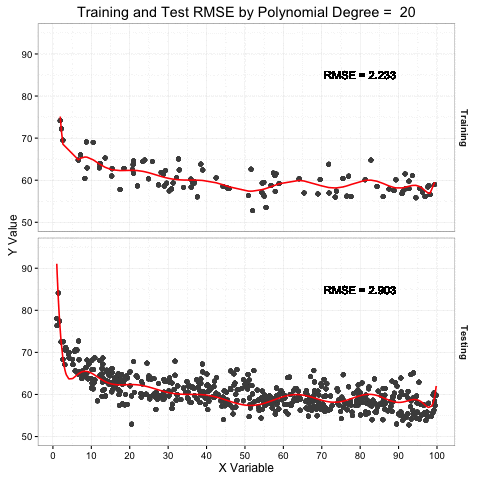
This is not the best example of the Simpson’s Paradox, but I think it is illustrative. I have a set of 150 points taken with a GPS, most taken with a very high accuracy RTK network and other with less accurate SBAS. However, I do not know which shots were by which method. My end calculation are showing a bias towards my elevations being about a half foot too high when compared to highly accurate LiDAR derived DEM. I began to suspect a vertical datum shift, but metadata and RTK information said they matched; then again, metadata and documentation can lie. I figured I could use the Vertical Dilution of Precision (VDOP) to detect the datum shift, so I set about plotting my data and estimating the linear trend.
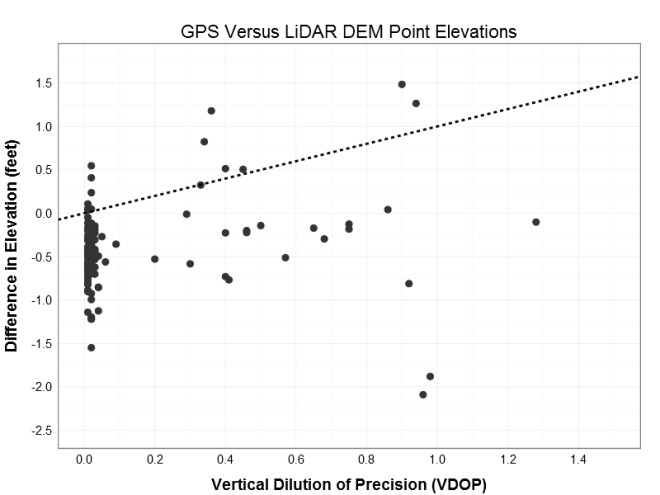
The slope of the linear fit is 0.44 suggesting a positive relationship between the VDOP and difference in elevation (increasing VDOP by 1 increases the delta elevation by 0.44 feet). However, this is counter intuitive given the increase in error (VDOP) is getting me closer to my target (delta elevation of zero). Albeit counterproductive, this suggests that the higher VDOP points are closer to target, and therefore may be the points in the correct vertical datum. based on a visual inspection of the data, I drew an arbitrary class boundary at VDOP = 0.25 to establish possible RTK vs. non-RTK groups. Given these the same treatment illustrates the point of this post: Simpson’s Paradox!
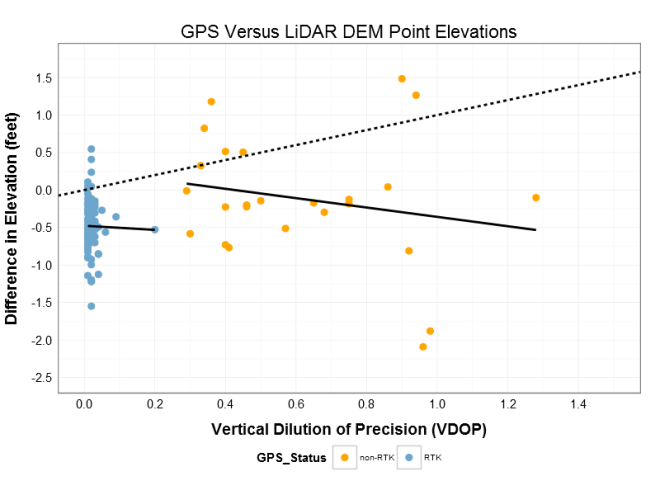
When the points are grouped and the linear trend is refit, the slope changes direction! Instead of the positive 0.44 of the aggregated data,the individual groups have negative slopes of -0.62 for non-RTK and -1.3 for RTK (the direction of the slope is important part here). Firstly, the negative slope for each group gives the more initiative result that higher VDOP error moves us further from the target elevation. Secondly, the possible RTK group is close to the 0.5 ft bias I had detected, while the non-RTK is closer to zero (which much more variability (see below) because of the lack of real-time post processing). The fact that the signs switched (the paradox part) and that my group means are close to the biased and true targets (see below) gives me confidence that I can identify the points with the incorrect vertical data and correct for it! Thank you Simpson’s Paradox.

PS – These are high accuracy GPS measurement, but they are used only for planning purposes; not construction of regulatory use. If that point at VDOP 0.2 should actually be in the non-RTK cluster and it gets correct as if it were RTK, no one will get hurt and no buildings will fall over.
*** below is the R code for the ggplot2 examples, but because it is a client project I cannot share the data.
Data prep and first plot
## Prepare data for plotting
pdat <- data.frame(elev_diff2 = sdat1$elev_diff, vdop = sdat1$vdop, hdop = sdat1$hdop, pdop = sdat1$pdop, Status = ifelse(sdat1$vdop > 0.25, "non-RTK", "RTK"))
colnames(pdat) <- c("Elevation_Difference", "VDOP", "HDOP", "PDOP", "GPS_Status")
## first plot
ggplot(data = pdat, aes(x = VDOP, y = Elevation_Difference)) +
geom_point(size = 5, shape = 20, color = "gray20") +
geom_abline(size = 1, linetype = "dashed") +
# geom_smooth(method="lm", se=FALSE) +
# stat_smooth(method="lm", formula=y~1, se=FALSE) +
# scale_color_manual(values = c("orange", "skyblue3")) +
scale_x_continuous(breaks = seq(0, 1.5, 0.2), limits = c(0, 1.5)) +
scale_y_continuous(breaks = seq(-2.5, 1.75, 0.5), limits = c(-2.5, 1.75)) +
ylab("Difference in Elevation (feet)") +
xlab("Vertical Dilution of Precision (VDOP)") +
ggtitle("GPS Versus LiDAR DEM Point Elevations") +
theme_bw() +
theme(
axis.title.y = element_text(hjust = 0.5, vjust = 1.5, angle = 90, size=rel(1), face="bold"),
axis.title.x = element_text(vjust = -0.5, size=rel(1), face="bold"),
title = element_text(size=rel(1.5)),
axis.ticks.x = element_blank(),
axis.ticks.y = element_blank(),
axis.text.x = element_text(size=rel(1.5)),
axis.text.y = element_text(size=rel(1.5)))
Second Plot
ggplot(data = pdat, aes(x = VDOP, y = Elevation_Difference ,
group = GPS_Status, color = GPS_Status)) +
geom_point(size = 5, shape = 20) +
geom_abline(size = 1, linetype = "dashed") +
geom_smooth(method="lm", se=FALSE, color = "black", size = 1) +
# stat_smooth(method="lm", formula=y~1, se=FALSE) +
scale_color_manual(values = c("orange", "skyblue3")) +
scale_x_continuous(breaks = seq(0, 1.5, 0.2), limits = c(0, 1.5)) +
scale_y_continuous(breaks = seq(-2.5, 1.75, 0.5), limits = c(-2.5, 1.75)) +
ylab("Difference in Elevation (feet)") +
xlab("Vertical Dilution of Precision (VDOP)") +
ggtitle("GPS Versus LiDAR DEM Point Elevations") +
theme_bw() +
theme(
axis.title.y = element_text(hjust = 0.5, vjust = 1.5, angle = 90, size=rel(1), face="bold"),
axis.title.x = element_text(vjust = -0.5, size=rel(1), face="bold"),
title = element_text(size=rel(1.5)),
axis.ticks.x = element_blank(),
axis.ticks.y = element_blank(),
legend.position = "bottom",
axis.text.x = element_text(size=rel(1.5)),
axis.text.y = element_text(size=rel(1.5)))
Third plot
ggplot(data = pdat, aes(x = VDOP, y = Elevation_Difference ,
group = GPS_Status, color = GPS_Status)) +
geom_point(size = 5, shape = 20) +
geom_abline(size = 1, linetype = "dashed") +
geom_smooth(method="lm", se=FALSE, color = "black", size = 1) +
stat_smooth(method="lm", formula=y~1, se=TRUE, size = 1) +
scale_color_manual(values = c("orange", "skyblue3")) +
scale_x_continuous(breaks = seq(0, 1.5, 0.2), limits = c(0, 1.5)) +
scale_y_continuous(breaks = seq(-2.5, 1.75, 0.5), limits = c(-2.5, 1.75)) +
ylab("Difference in Elevation (feet)") +
xlab("Vertical Dilution of Precision (VDOP)") +
ggtitle("GPS Versus LiDAR DEM Point Elevations") +
theme_bw() +
theme(
axis.title.y = element_text(hjust = 0.5, vjust = 1.5, angle = 90, size=rel(1), face="bold"),
axis.title.x = element_text(vjust = -0.5, size=rel(1), face="bold"),
title = element_text(size=rel(1.5)),
axis.ticks.x = element_blank(),
axis.ticks.y = element_blank(),
legend.position = "bottom",
axis.text.x = element_text(size=rel(1.5)),
axis.text.y = element_text(size=rel(1.5)))
Linear fits
m1 <- lm(elev_diff ~ vdop, data = sdat1)
summary(m1)
m2 <- lm(elev_diff ~ vdop, data = sdat1[which(sdat1$vdop > 0.25),])
summary(m2)
m3 <- lm(elev_diff ~ vdop, data = sdat1[which(sdat1$vdop < 0.25),])
summary(m3)